Understanding MRI image harmonization
Magnetic resonance imaging (MRI) is a valuable and versatile medical imaging method, yet it often displays heterogeneous image quality across different protocols and scanners.
We regularly see image variations in data analysis, particularly when handling real-world data from multiple imaging centers. These discrepancies can stem from different scanner manufacturers, scanner models or acquisition protocols. This leads to notable differences in image intensity values. Such variations present substantial challenges in creating universally applicable and reliable models1. They also impact the reproducibility of quantitative imaging biomarkers and radiomics features2, crucial for developing clinical decision support systems. This variability is particularly prominent in Magnetic Resonance (MR) images because, unlike Computed Tomography (CT) scans, where intensity values are standardized to Hounsfield Units (which also require certain degree of harmonization), MR image intensities are not normalized to a standard unit of measurement. As can be observed in Figure 1, MRI scans acquired with different echo times (TE) and repetition times (TR) result in differences in image intensities.

Efforts to standardize MRI scans often encounter challenges due to variations in hardware, software, or scanning settings which can impact the accuracy of quantitative results. This issue becomes particularly important when using advanced technologies like Artificial Intelligence (AI) or digital analysis methods. In multi-site or long-term studies, where consistency is critical, these variations can significantly affect the reliability of the findings. Therefore, addressing these challenges is crucial to ensure the quality and consistency of MRI data in such research.
From traditional methods to Deep Learning techniques that manage variations in medical imaging
Traditional computer vision techniques often employ methods to normalize, center, and standardize image intensities. These techniques typically scale the intensity values to a common range, adjust the mean intensity to zero, and ensure unit variance. However, these conventional methods have yet to achieve robust generalization performance in medical imaging, mainly when working with data from various imaging centers.
Addressing these challenges necessitates the incorporation of data from multiple imaging centers into the training process. However, collecting such diverse data is challenging due to the need for substantial information from these centers. While tradicional denoising algorithms such as Diffusion Anisotropic, N4, or bias field correction filters have been employed to enhance image quality3,4,5, their capacity to generalize across different noise sources and the substantial computational cost involved restricts their practical application in clinical settings.
To overcome the limitations of traditional methods, exploring Deep Learning techniques to manage variations in medical imaging is crucial. These techniques focus on developing image standardization methods capable of handling the heterogeneities observed in the field. Image generation technique, which create new images usually based on existing ones6 have become increasingly relevant in medical imaging due to the generalization challenges AI-based solutions face.
Recent years have seen exciting advancements in this domain, with notable achievements in tasks like image inpainting (filling in missing parts of images)7, enhancing image quality or resolution8,9, noise reduction10,11,12, and style transfer using techniques like Cycle-GAN13,14.
Quibim’s methodology
On this regard, Quibim has introduced a novel methodology that leverages self-supervised techniques and makes use of the frequency domain in an unique manner, to synthetically generate contrast variations from original, high-quality, and homogeneous images that act as the reference group. This reference group reunites the most desired image feature representations to which the rest of the images will imitate, and is only composed by the best 13 cases, as thousands of alterations are obtained for each case. This approach involves training a Convolutional Neural Network (CNN) to standardize these varied generated contrasts to the same reference group. This simple yet innovative solution can be generalized effectively, particularly as the network is exposed to a wide range of contrast alterations for learning purposes.
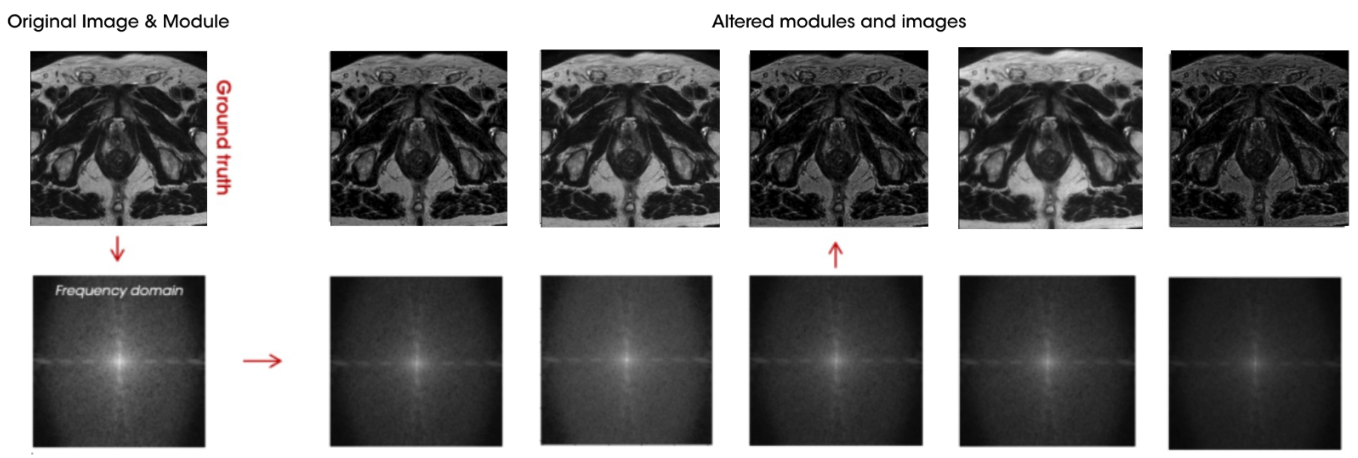
Figure 2 shows an example of alterations performed on a sample from the reference group. The reference image is transformed into the frequency domain where thousands of alterations are applied (bottom row). Finally, these altered frequency domain images are transformed back to the imaging domain where the synthetically alterations on image intensities are observed (top row).
Working within the frequency domain has been the key aspect of this methodology, data in K-space is a fundamental concept in MRI, representing the raw data acquired during an MRI scan. Data in K-space, obtained through magnetic gradients and radiofrequency pulses, are transformed via Fourier transformation from the frequency domain to the imaging domain, resulting in an interpretable image. K-space can be conceptualized as a matrix with points representing specific frequencies and phases of the signal, where the center captures low-frequency components, and the periphery captures high-frequency components, according to this, it is posible to perform changes in subtle features such as contrast or brightness of the image while keeping the content of the original sample. Original K space raw data is no longer available when the image reaches the PACS, but it is possible to access to the frequency domain by performing the Fourier transformation.
This approach involves training a Convolutional Neural Network (CNN) to standardize these varied generated contrasts to the same reference group. This simple yet innovative solution can be generalized effectively, particularly as the network is exposed to a wide range of contrast alterations for learning purposes, allowing.paired training, aligning each sample with its altered counterpart, thus enhancing the robustness and adaptability of the model across diverse medical imaging scenarios.
Figure 3 shows some results of our innovative solution, we can see how we can standardize image intensities across a dataset with an heterogenoeus MRI dynamic range.
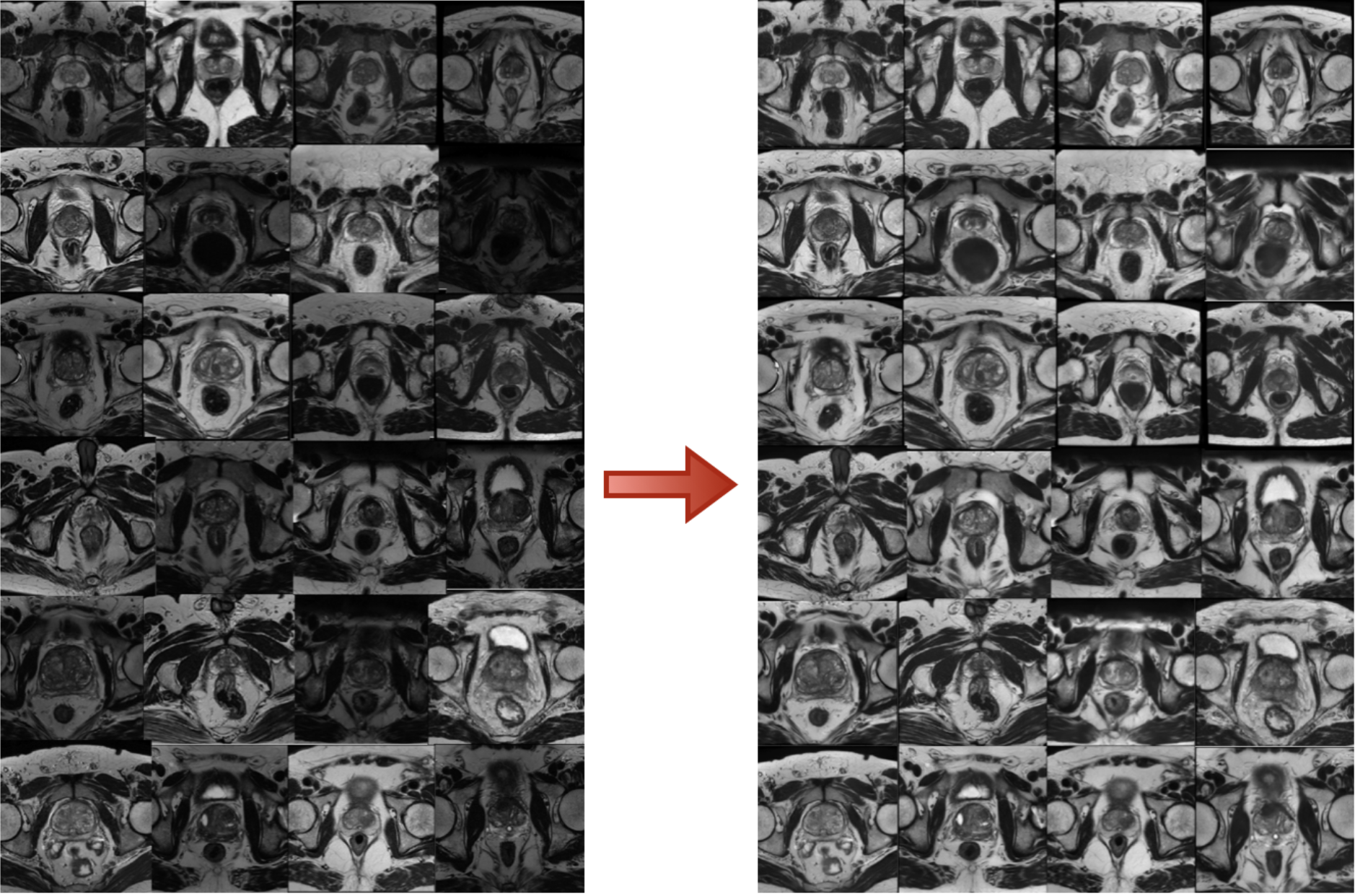
Quibim’s approach marks a significant movement forward in overcoming the longstanding challenges of image standardization in medical imaging, helping to find more reliable and accurate diagnostic tools.
References
- Kushol, R., Parnianpour, P., Wilman, A.H. et al. Effects of MRI scanner manufacturers in classification tasks with deep learning models. Sci Rep 13, 16791 (2023). https://doi.org/10.1038/s41598-023-43715-5
- Mi H, Yuan M, Suo S, Cheng J, Li S, Duan S, Lu Q. Impact of different scanners and acquisition parameters on robustness of MR radiomics features based on women’s cervix. Sci Rep. 2020 Nov 23;10(1):20407. doi: 10.1038/s41598-020-76989-0. PMID: 33230228; PMCID: PMC7684312.
- Caio A. Palma, Fábio A.M. Cappabianco, Jaime S. Ide, Paulo A.V. Miranda, Anisotropic Diffusion Filtering Operation and Limitations – Magnetic Resonance Imaging Evaluation, IFAC Proceedings Volumes,
- Tustison NJ, Avants BB, Cook PA, Zheng Y, Egan A, Yushkevich PA, Gee JC. N4ITK: improved N3 bias correction. IEEE Trans Med Imaging. 2010 Jun;29(6):1310-20. doi: 10.1109/TMI.2010.2046908. Epub 2010 Apr 8. PMID: 20378467; PMCID: PMC3071855.
- SONG, Shuang; ZHENG, Yuanjie; HE, Yunlong. A review of Methods for Bias Correction in Medical Images. Biomedical Engineering Review, [S.l.], v. 1, n. 1, sep. 2017. ISSN 2375-9151.
- Roberto Gozalo-Brizuela, Eduardo C. Garrido-Merchán. A survey of Generative AI Applications, may. 2023
- Hanyu Xiang, Qin Zou, Muhammad Ali Nawaz, Xianfeng Huang, Fan Zhang, Hongkai Yu, Deep learning for image inpainting: A survey, Pattern Recognition, Volume 134, 2023,109046,ISSN 0031 3203,https://doi.org/10.1016/j.patcog.2022.109046.
- Yamashita, Koki & Markov, Konstantin. (2020). Medical Image Enhancement Using Super Resolution Methods. 10.1007/978-3-030-50426-7_37.
- Yuan Ma, Kewen Liu, Hongxia Xiong, Panpan Fang, Xiaojun Li, Yalei Chen, Zejun Yan, Zhijun Zhou, Chaoyang Liu, Medical image super-resolution using a relativistic average generative adversarial network, Nuclear Instruments and Methods in Physics Research Section A: Accelerators, Spectrometers, Detectors and Associated Equipment, Volume 992, 2021, 165053, ISSN 0168-9002, https://doi.org/10.1016/j.nima.2021.165053.
- Kaur P, Singh G, Kaur P. A Review of Denoising Medical Images Using Machine Learning Approaches. Curr Med Imaging Rev. 2018 Oct;14(5):675-685. doi: 10.2174/1573405613666170428154156. PMID: 30532667; PMCID: PMC6225344.
- Arshaghi, Ali & Ashourian, Mohsen & Ghabeli, Leila. (2020). Denoising Medical Images Using Machine Learning, Deep Learning Approaches: A Survey. Current Medical Imaging Formerly Current Medical Imaging Reviews. 16.10.2174/1573405616666201118122908.
- Junshen Xu, Elfar Adalsteinsson, Deformed2Self: Self-Supervised Denoising for Dynamic Medical imaging arXiv:2106.12175
- Joonhyeok Yoon, Sooyeon Ji, Eun-Jung Choi, Hwihun Jeong, and Jongho Lee, Seoul National University, Seoul, Korea, Republic of, Paired CycleGAN-based Cross Vendor Harmonization https://archive.ismrm.org/2022/1881.html
- Modanwal, Gourav & Vellal, Adithya & Buda, Mateusz & Mazurowski, Maciej. (2020). MRI image harmonization using cycle-consistent generative adversarial network. 36. 10.1117/12.2551301.
